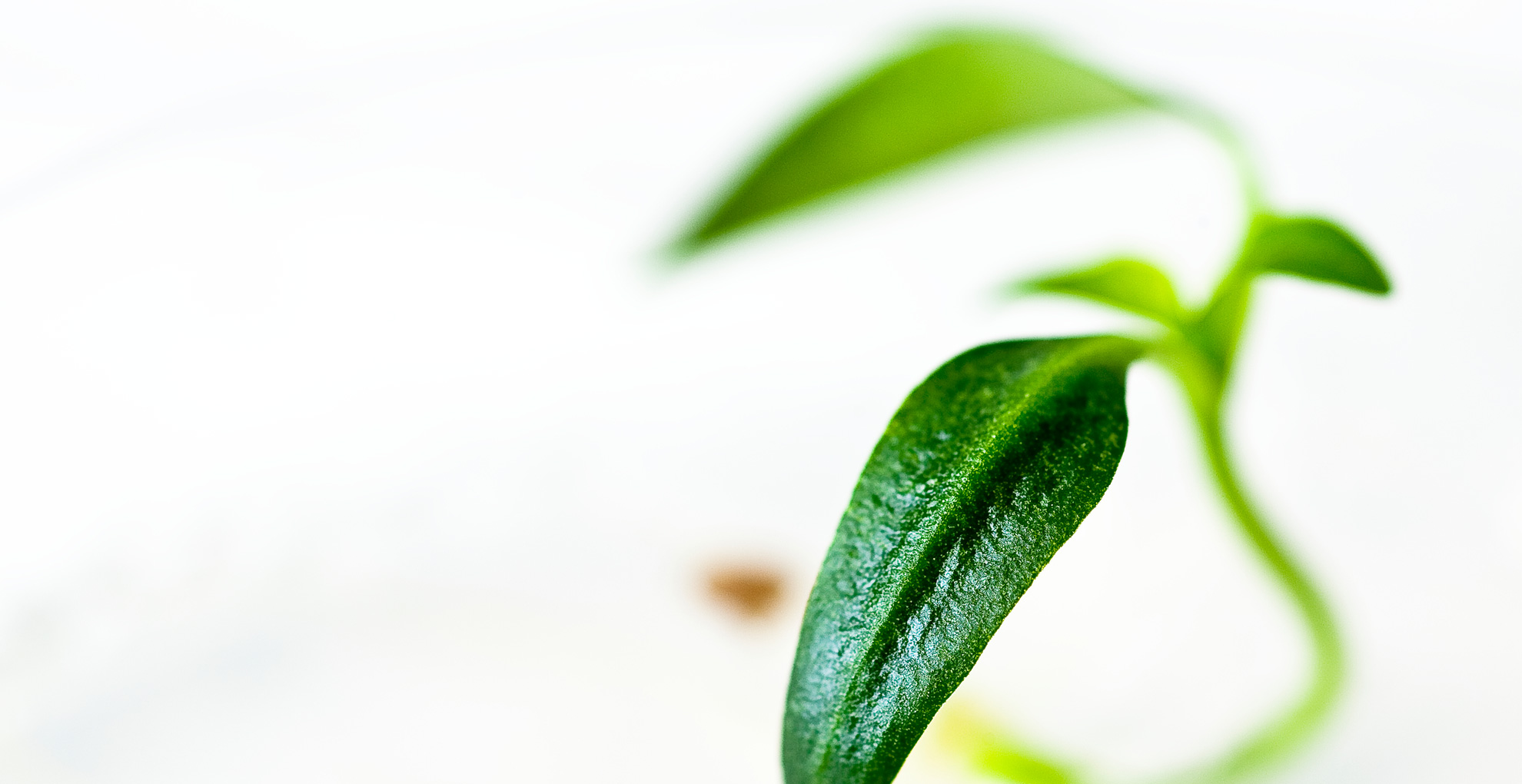

Creativity in supporting in silico plantbreeding
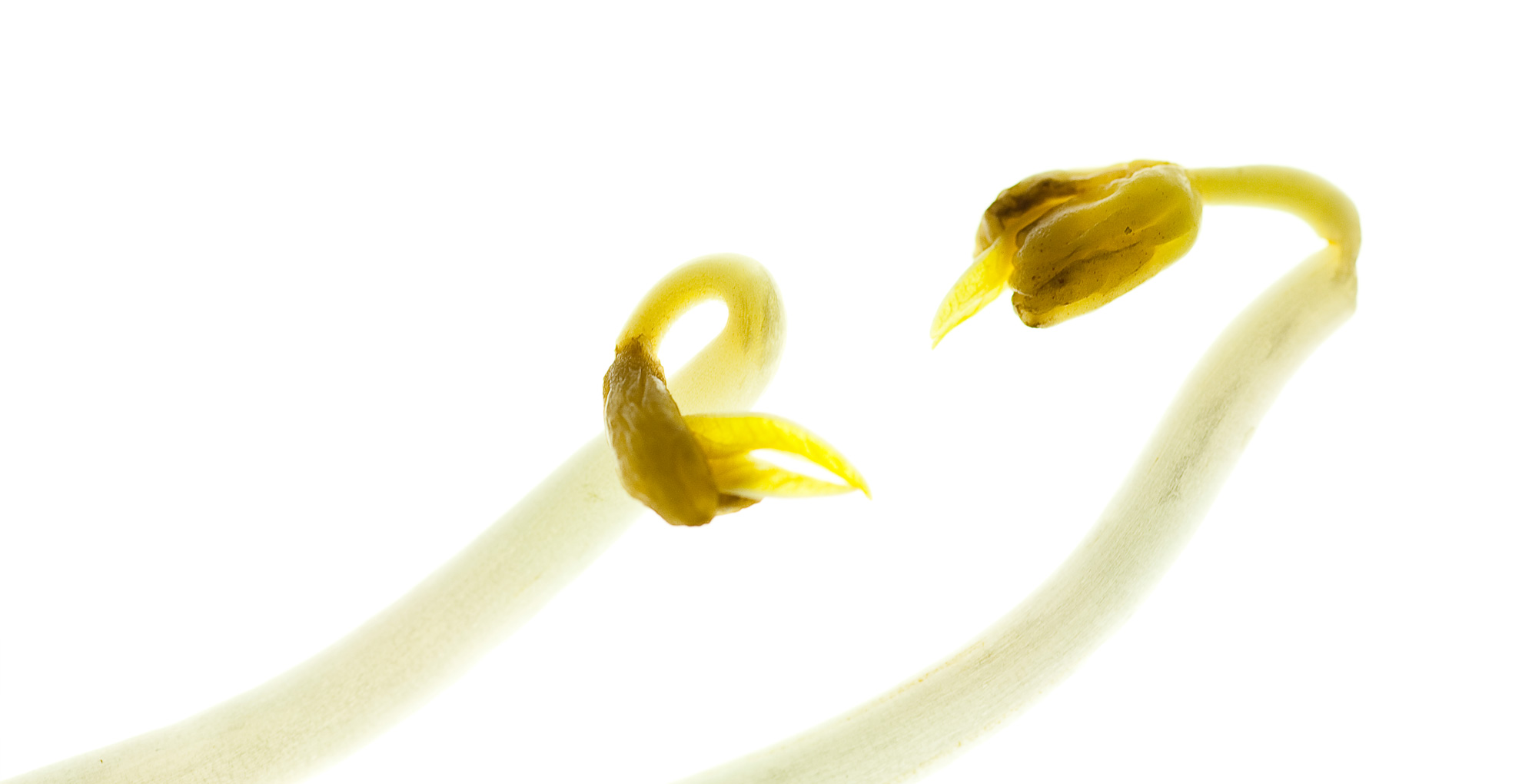

Virtual Laboratory Plant Breeding
The VLPB is:
An association of 6 plant breeding companies that initiates projects in the domains of e-bioscience, bioinformatics and IT-infrastructure in which the plant breeding industry and associated knowledge institutes collaborate.
VLPB activities are of pre-competitive nature. The VLPB is active in the areas of research technology and infrastructure and herewith promotes and speeds up innovation in the plant-breeding industry.
The VLPB aims to:
- To secure real-life use of VLPB methods and infrastructure by end-users via a demand-driven development strategy.
- To create and maintain the requirements for a vibrant user community.
- To scout and to implement novel developments in methodology and technology.
- To secure long term commitment and continuity.
Meet the members:
VLPB members are plant breeding companies and companies active in the field of plant breeding. Next to members the VLPB count a number of associated members. Associated members are from academia and public knowledge institutions.
Members






Associated Members



VLPB board:
Board
Laurens Kroon – Bejo zaden
Marcel Reinders – TUDelft
Simon Langeveld – East-West Seed
Richard Visser – Wageningen University
Robert Graveland (chairman) – HZPC
Our Latest Projects
VLPB completed and running projects by 2020
VLPB is continuously initiating new projects and activities on the basis of the prioritised needs of the members. The VLPB members establish annually a budget which enables to respond quickly and agile to novel developments in – and application of technology. In practice this results in funding of about 5 small projects with a size between 1,5k€ and 20k€. Budget for small projects is allocated in February of each year on basis of ranking of small projects which have been submitted by the end of the previous year.
On basis of common ground and joint commitment VLPB members also initiate large projects in collaboration with Dutch Universities. Finally VLPB members may also initiate projects dedicated to development and implementation of specific software. Such projects are generally executed by professional software providers.

Ultra-long read 3rd-generation DNA sequencing
The complete genomes of two economically relevant plant species; tomato and potato were sequenced using

TKI-U Discovery toolkit for small-RNAs relevant to plant breeding
This project aimed to guide plant breeding researchers onto the path of small RNA discovery/analysis

RepeatDB
Repeat regions are important sequences within a chrosomome. They play roles in regulation, genome structure,
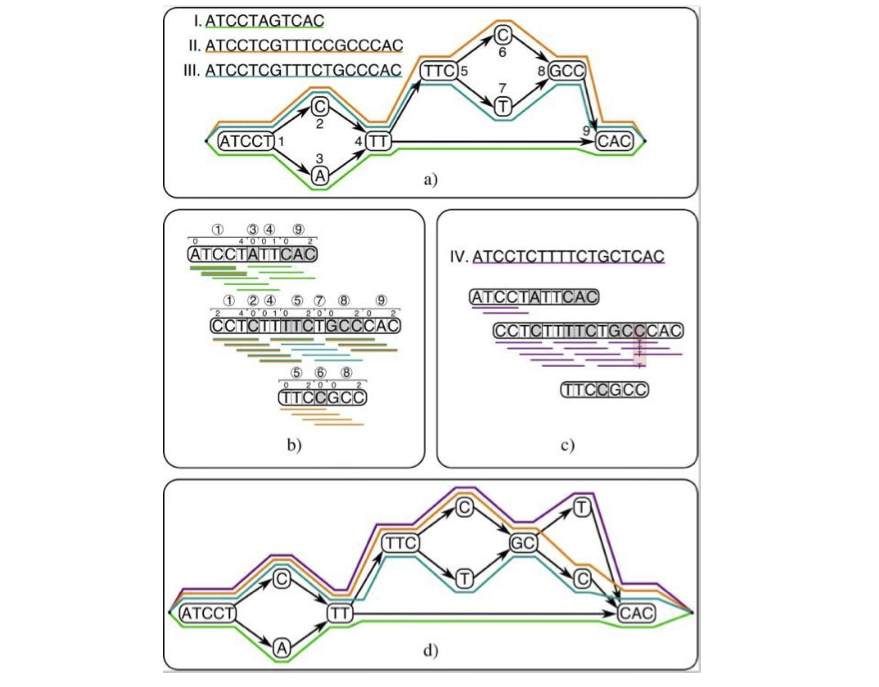
Pangenome read alignment and variant calling
Pan-genome references, also known as graph reference genomes, can represent multiple strains in a single
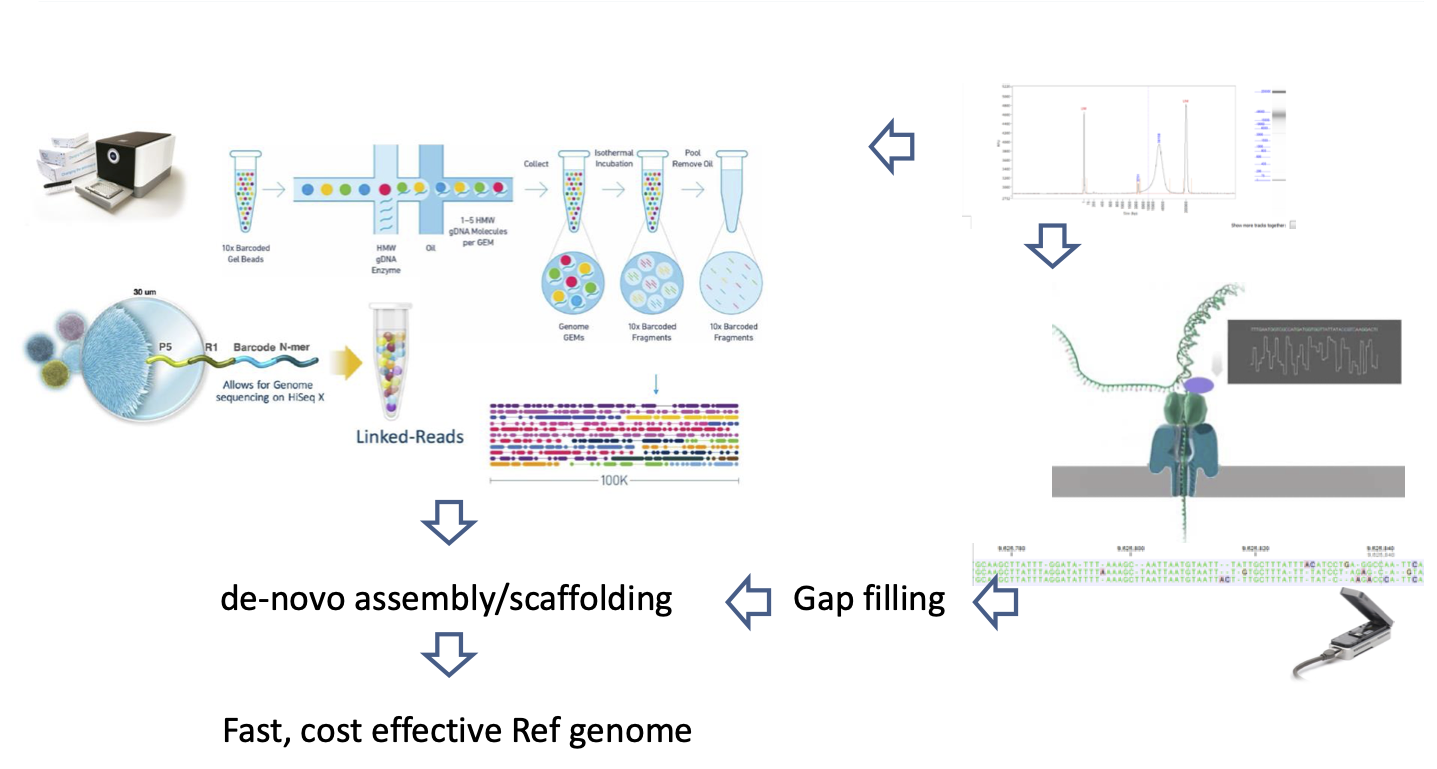
Improving genome assemblies by combined 10XGenomics and nanopore technology
This project’s aim was to generate an improved reference genome of potato (Solanum tuberosum Phureja
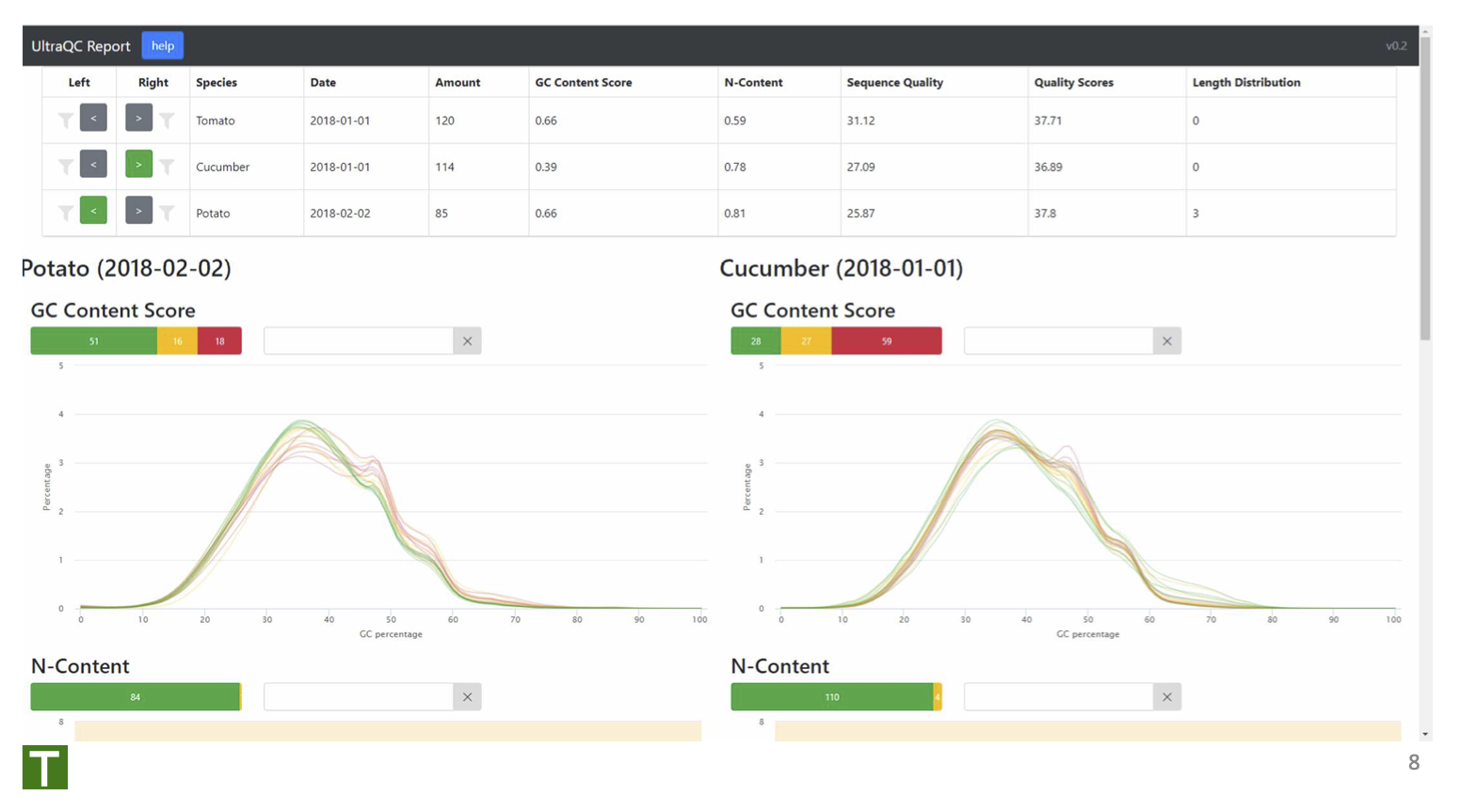
High-throughput quality assessment of sequencing data (student project)
The aim of this project was to provide key summary statistics from whole genome DNA

Haplotype reconstruction algorithms
Long-read sequencing enables for the first time to reconstruct chromosome length haplotypes. We have recently
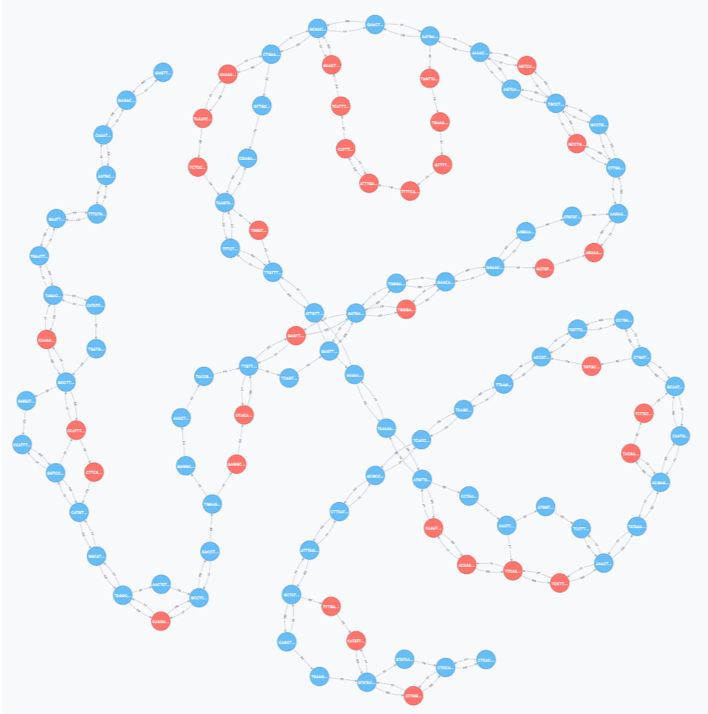

Exploring pan-genomics for crops
The aim was to evaluate how pan-genomics can facilitate data management and ease specific data
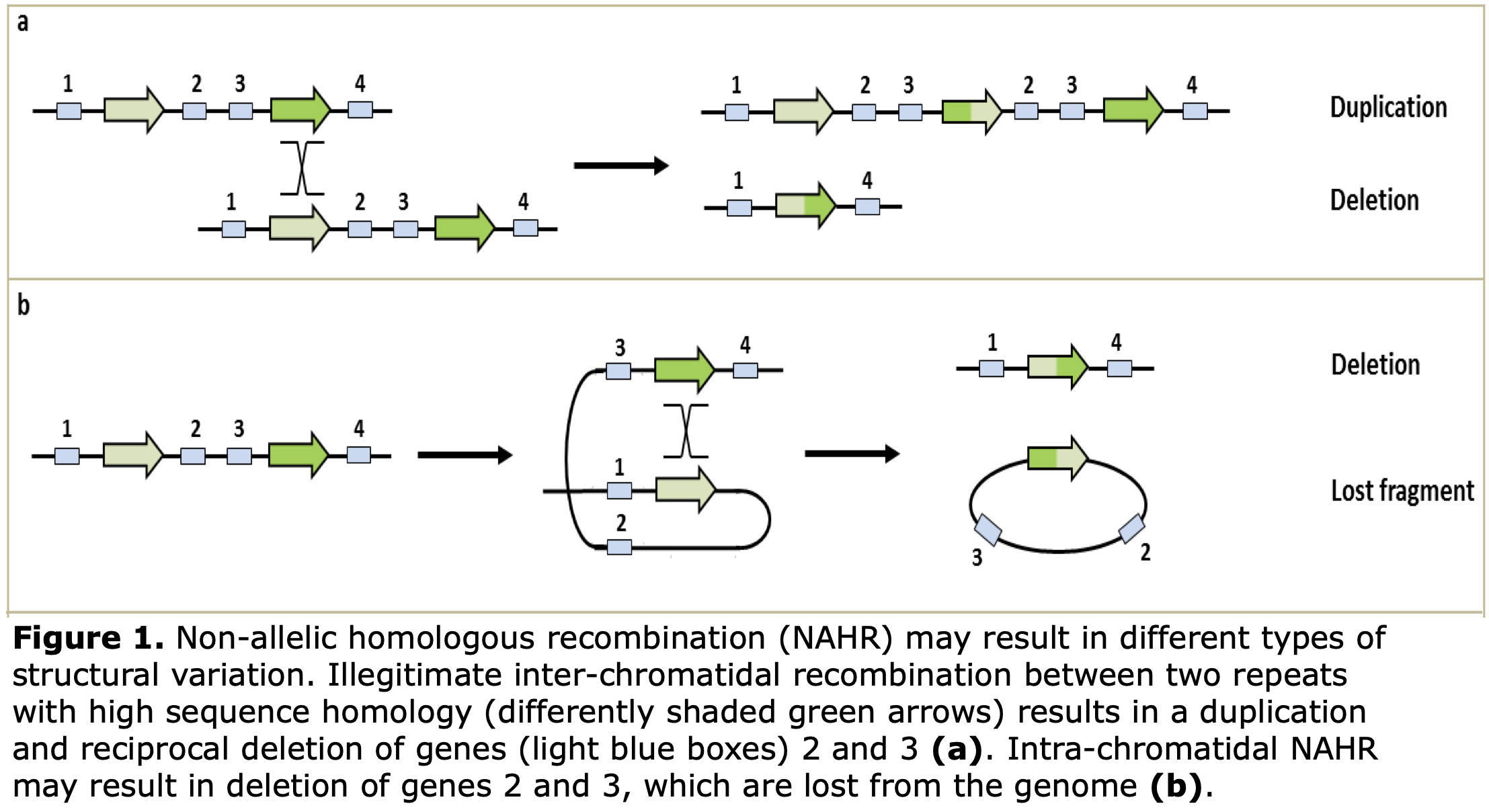
Copy Number Variation
The key challenge is to reliably detect CNV on (very) low coverage datasets, to link
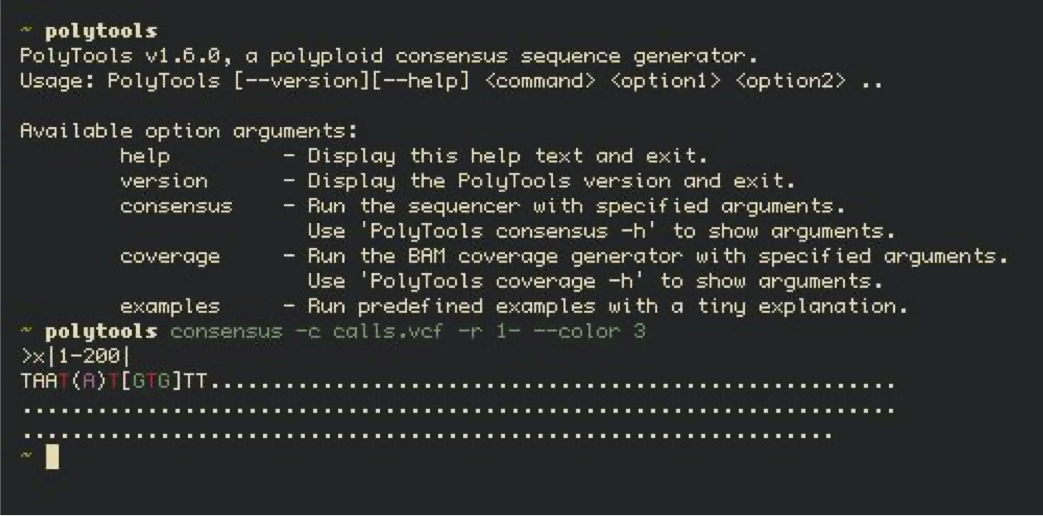
Consensus sequence generator (student project)
The objective of the consensus sequence generator was to develop a command-line tool that allows
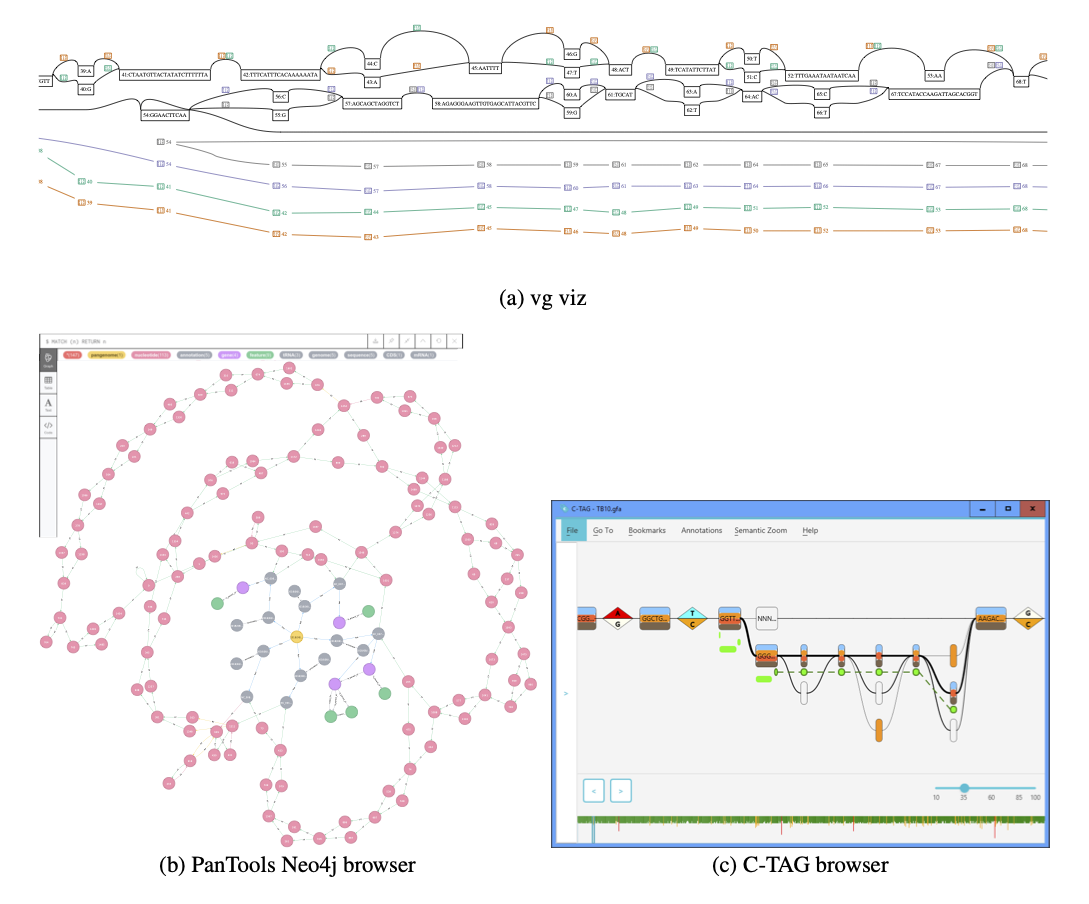
Comparison of pan-genome approaches (student project)
What are the functionalities, limitations, strengths and weaknesses of various pan-genome representations?The aim was to
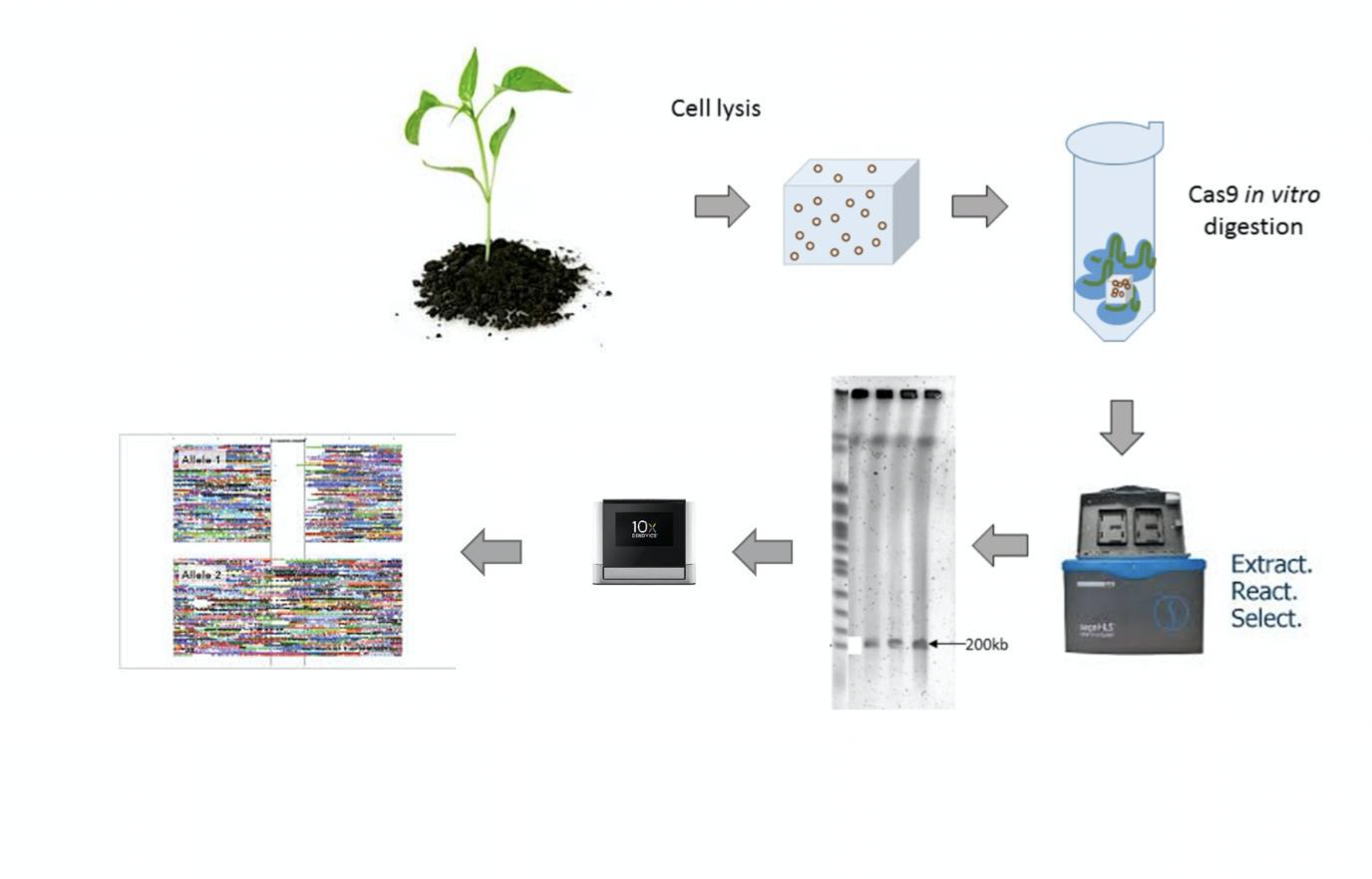
Building the Green Hapmap, 10X Genomics
A promising technology to deal with haplotype assembly in a cost-effective manner is 10x Genomics.
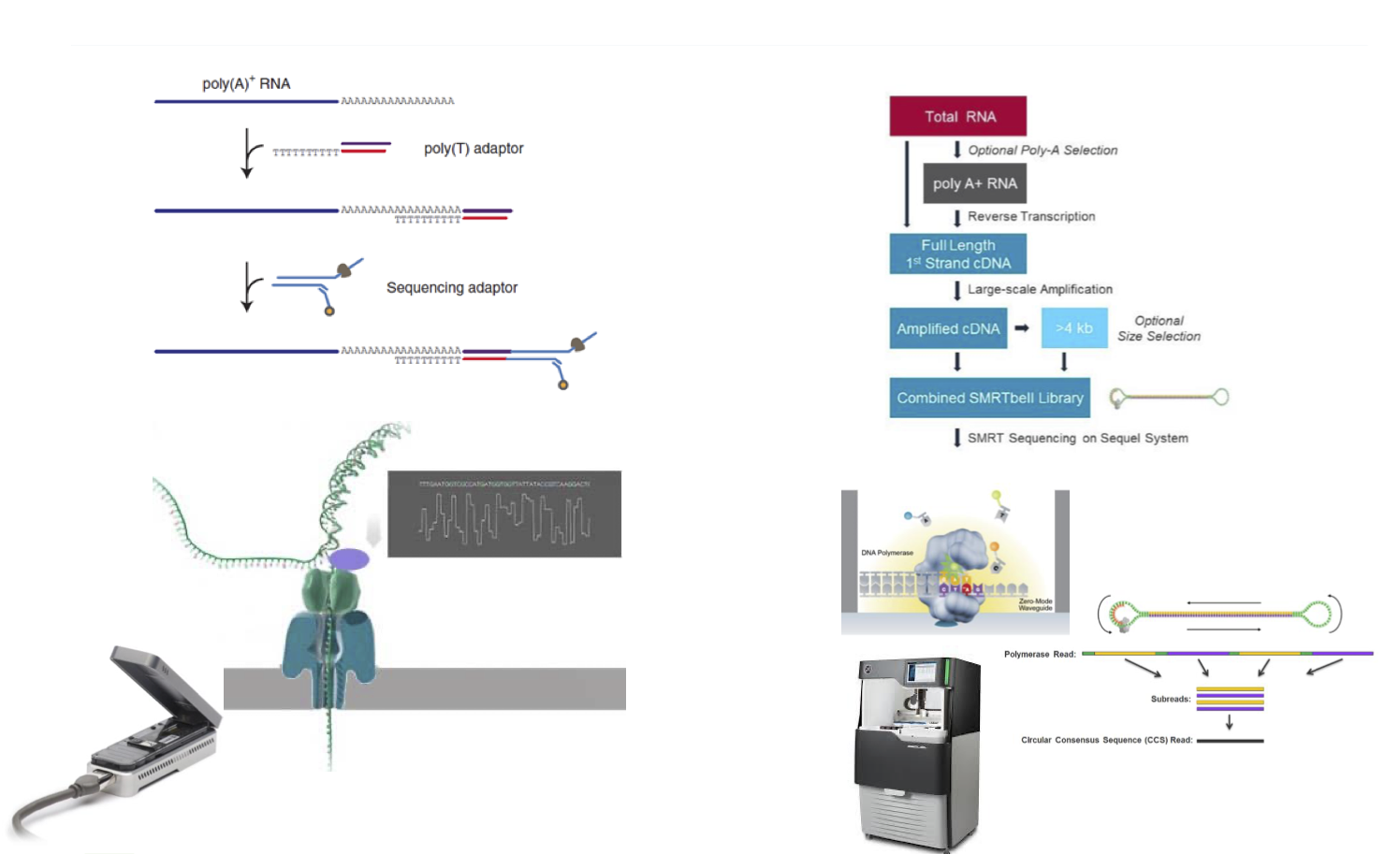
Benchmarking Crop Direct RNA sequencing versus full length IsoSeq
The aim was to explore technological possibilities for direct RNA sequencing using nanopore technology compared

Application of candidate gene prioritization to QTL data
In this project we used gene function prediction and functional overrepresentation in QTL regions to
